
WHY THIS MATTERS IN BRIEF
Being able to do things at the nanoscale, such as create molecular assemblers, means we need to be able to move things at the nanoscale.
 Interested in the Exponential Future? Connect, download a free E-Book, watch a keynote, or browse my blog.
Interested in the Exponential Future? Connect, download a free E-Book, watch a keynote, or browse my blog.
Today we’re building lots of nanoscale things, like cancer killing nanobots, bio-hybrid robots that are part plant, part robot, and all manner of other nanobots, nanomachines and molecular robots that are being used to create, among other things, the world’s first molecular assemblers. And asides from being tiny they all have one thing in common – they all need engines to move. Recently I’ve talked about the development of new enzyme engines and molecular motors that could power some of these nanoscale bots, and now researchers have announced they’ve created a DNA motor that’s also the world’s fastest nanoscale motor that uses RNA as its fuel source.
“Nanoscale motors have tremendous potential for applications in biosensing, in building synthetic cells and also for molecular robotics,” says Khalid Salaita, a senior author of the paper and a professor of chemistry at Emory University School of Medicine. “DNA origami allowed us to tinker with the structure of the motor and tease out the design parameters that control its properties.”
The new DNA motor is rod-shaped and uses RNA fuel to roll persistently in a straight line, without human intervention, at speeds up to 100 nanometers per minute. That’s up to 10 times faster than previous DNA motors.
“Our engineered DNA motor is fast,” Ke says, “but we still have a long way to go to achieve the versatility and efficiency of nature’s biological motors. Ultimately, the goal is to make artificial motors that match the sophistication and functionality of proteins that move cargo around in cells and allow them to perform various functions.”
Making things out of DNA, nicknamed DNA origami after the traditional Japanese paper folding craft, takes advantage of the natural affinity for the DNA bases A, G, C and T to pair up with one another. By moving around the sequence of letters on the strands, researchers can get the DNA strands to bind together in ways that create different shapes. The stiffness of DNA origami can also easily be adjusted, so they remain straight as a piece of dry spaghetti or bend and coil like boiled spaghetti.
Growing computational power, and the use of DNA self-assembly for the genomics industry, have greatly advanced the field of DNA origami in recent decades. Potential uses for DNA motors include drug delivery devices in the form of nanocapsules that open up when they reach a target site, nanocomputers and nanbots working on nanoscale assembly lines – something that’s already been demonstrated after scientists elsewhere created what can ostensibly be thought of as the world’s first molecular assembler.
“These applications may seem like science fiction now, but our work is helping move them closer to reality,” says Alisina Bazrafshan, an Emory PhD candidate and first author of the new paper.
One of the biggest challenges of DNA motors is the fact that rules governing motion at the nanoscale are different than those for objects that humans can see. Molecular-scale devices must fight their way through a constant barrage of molecules. These forces can cause such tiny devices to drift randomly like grains of pollen floating on the surface of a river, a phenomenon known as Brownian motion.
The viscosity of liquids also makes a much larger impact on something as tiny as a molecule, so water becomes more like molasses.
Many prior DNA motors “walk” with a mechanical leg-over-leg motion. The problem is that two-legged versions tend to be inherently unstable. Walking motors with more than two legs gain stability but the extra legs slow them down.
The Emory researchers solved these problems by designing a rod-shaped DNA motor that rolls. The rod, or “chassis” of the motor consists of 16 DNA strands bound together in a four-by-four stack to form a beam with four flat sides. Thirty-six bits of DNA protrude from each face of the rod, like little feet.
To fuel its motion, the motor is placed on a track of RNA, a nucleic acid with base pairs that are complementary to DNA base pairs. The RNA pulls at the DNA feet on one face of the motor and binds them to the track. An enzyme that targets only RNA that is bound to DNA then quickly destroys the bound RNA. That causes the motor to roll, as the DNA feet on the next face of the motor get pulled forward by their attraction to RNA.
The rolling DNA motor forges a persistent path, so it continues to move in a straight line, as opposed to the more random motion of walking DNA motors. The rolling motion also adds to the new DNA motor’s speed: It can travel the length of a human stem cell within two or three hours. Previous DNA motors would need about a day to cover that same distance, and most lack the persistence to make it that far.
One of the biggest challenges was measuring the speed of the motor at the nanoscale. That problem was solved by adding fluorescent tags on either end of the DNA motor and optimizing imaging conditions on a fluorescent microscope.
Through trial and error, the researchers determined that a stiff rod shape was optimal for moving in a straight line and that 36 feet on each face of the motor provided optimal density for speed.
“We provided a tunable platform for DNA origami motors that other researchers can use to design, test and optimize motors to further advance the field,” Bazrafshan says. “Our system allows you to test the effects of all kinds of variables, such as chassis shape and rigidity and the number and density of legs to fine tune your design.”
For instance, what variables would give rise to a DNA motor that moves in circles? Or a motor that turns to go around barriers? Or one that turns in response to a particular target?
“We hope other researchers will come up with other creative designs based on these findings,” Bazrafshan says.
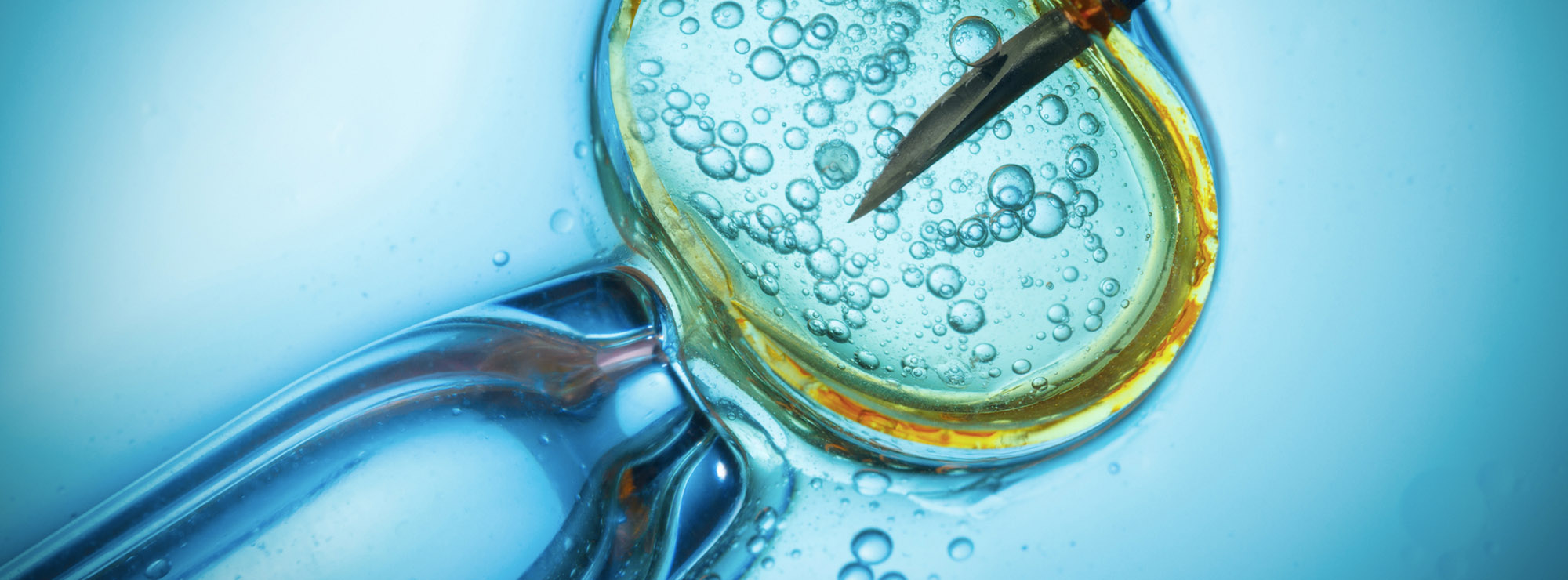














sounds a little like the Matrix babies to me