WHY THIS MATTERS IN BRIEF
With so much information being generated we need a new way to store it all, and DNA is a perfect storage solution.
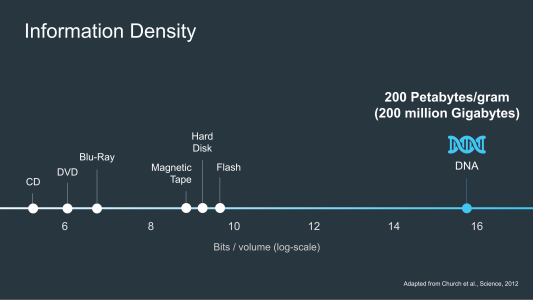
In a world awash with data, DNA is a hugely efficient and compact way to store it. After all, the data on every mobile, PC, and server rack on the planet could fit in a shoebox if it was stored in DNA and not on flash drives, hard drives or tape like it is today, and just recently scientists managed to cram an amazing 215 petabytes of information onto just one gram of DNA. DNA also has another advantage over these other storage mediums – it’s incredibly durable, DNA can last for hundreds of thousands of years without degrading so long, that is, as you keep it relatively cool and dry.
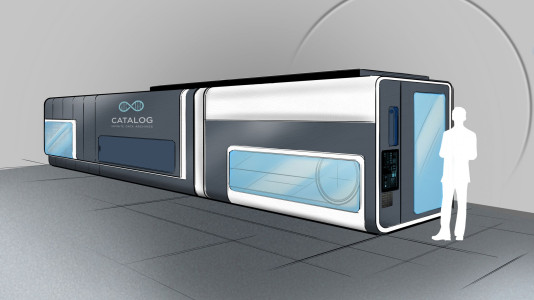
Now, with Microsoft announcing the roll out of its first cloud based DNA storage service starting in, or around, 2020, one of the fledgling DNA storage industry’s other venerable startups has unveiled plans for its prototype storage device – a hulking school bus sized machine that could one day convert movies or data archives into invisible pellets of DNA. The device is being built by Catalog DNA, who’ve I’ve discussed before, along with UK firm Cambridge Consultants.
Catalog claims its system is less costly than other DNA storage alternatives too. Instead of synthesizing unique strands of DNA, their process involves combining inexpensive, short, premade DNA strands into longer bits of DNA that carry and encode information.
Hyunjun Park, CEO and cofounder of Catalog, compares the process to the way metal letters are combined together to form words on an old fashioned printing press.
The companies provided a glimpse of their machine to MIT Technology Review, including a rendering, of what the walk-in lab will look like and a photograph of engineers outfitting a prototype, both of which are shown above.
According to Park, a single prototype machine will be completed early next year and will be able to write one terabit of data into DNA per day. That’s about as much data as fits on a laptop.
“Still not enough, but it is larger than what has ever been done before,” he says.
An actual commercial system, a single machine or a group of them able to store a petabit of data per day, he says, won’t be ready until 2021.
And let’s face it, this thing is huge. It’s no flash drive. The rendering shows a door and room enough inside for a couple of technicians. Inside there will need to be a hundred bags or bottles of ready-made DNA, and then an automated laboratory to mix the strands together and perform billions of reactions. You’ll also have to squeeze in a DNA sequencing machine, maybe a couple of them, to retrieve the data. The rendering has odes of ENIAC, the early computer built in the 1940s at the University of Pennsylvania, which glowed with over 18,000 vacuum tubes and filled a large room, about it.
“This is the first ever one. It can shrink, but we haven’t taken that challenge on the nose yet,” says Richard Hammond, head of synthetic biology at Cambridge Consultants, which takes on custom engineering projects.
Catalog, which raised $9 million in venture funds this summer, will not be selling the machines at first. Instead, when the single prototype device is finished, the company plans to allow partners to try out storing files in DNA as a service, although Park doesn’t say if any have signed up already.
Because it takes so long to turn bits into DNA and to get the information back out, don’t expect DNA data storage on your phone for a long time though. Rather, Park says that in the short to medium term at least the technology could replace long term archival storage on magnetic tapes.
Catalog has also been secretive about its approach, leading other scientists to say they can’t judge whether it makes sense. Victor Zhirnov at the Semiconductor Research Corporation in Durham, North Carolina, which is tracking developments in DNA storage, says the firm’s “library” idea is economically viable, in theory.
“By doing this they don’t need to synthesize new DNA for every new piece of information to store, instead they just have to remix their prefabricated DNA,” he adds.
As I mentioned earlier though Catalog isn’t the only firm hoping to scale up DNA storage. Luis Ceze, at the University of Washington, is collaborating with Microsoft, which also has plans for its own commercial DNA storage system, and who also want to automate the process. And both groups are in the running for funds from IARPA, the research organization of US intelligence agencies, which earlier this year said that it wants to use the latest in biological, DNA and molecular computing and storage technologies to collapse their hyperscale datacentre estate into something the size of an average office desk.
Several teams have also already shown it’s possible to store and retrieve information in the DNA of living bacteria, such as movies, and replay them, and elsewhere there are efforts afoot to turn everything from bacteria to even us humans into DNA, or biological, computers, with some tangible successes.
The problem is that converting bits into the As, Gs, Cs, and Ts of the genetic code is slow, and it’s a laborious process to retrieve the data. The cost of manufacturing customized DNA is also high, running near a million dollars to store a couple of high-resolution DVDs, so there’s a lot of work to be done before these futuristic technologies can become mainstream.